In neuroscience and epileptology research, one of the most persistent hurdles is the precise localization of intracranial electrodes. When working with post-implantation MRI or CT scans, we need a way to bridge the gap between raw medical imaging and actionable data. To address this, I developed PyLocator, an open-source Python tool designed to make the mapping of electrodes within MRI volumes more intuitive and reproducible.
https://pylocator.thorstenkranz.de/
The Problem: Spatial Mapping
When a patient undergoes electrode implantation, the resulting images show the sensors as high-intensity voids or signals within the brain. However, simply "seeing" them on a scan isn't enough for data analysis. We need their exact coordinates in a 3D space to correlate neural activity with specific anatomical structures.
I built PyLocator to streamline this transition—moving from a stack of 2D slices to a labeled 3D coordinate system.
How PyLocator Works
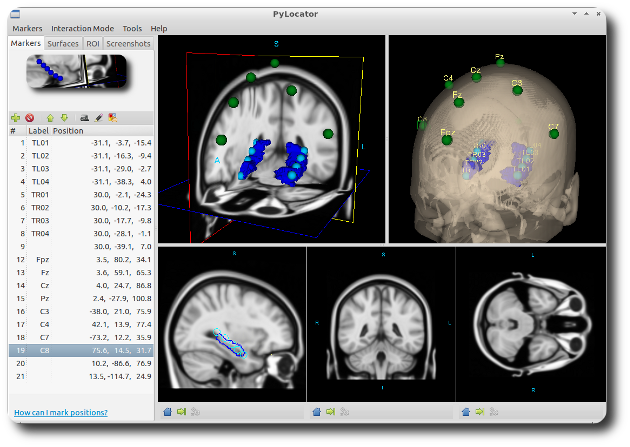
The core of the application is built on the Visualization Toolkit (VTK). It provides a synchronized interface where you can view MRI data through four different lenses simultaneously: axial, sagittal, and coronal slices, alongside a 3D "Planes" widget.
1. Marker Placement and Synchronization The workflow is centered on a marker system. By navigating the slices, you can place a 3D sphere (a "Marker") at the center of an electrode contact. Because all views are linked, moving a marker in the axial view instantly updates its position in the 3D rendering. This cross-referencing is vital for ensuring that a contact isn’t just "near" a structure, but precisely where it belongs.
2. Anatomical Context To provide better spatial orientation, PyLocator supports iso-surface rendering. This allows you to generate a 3D shell of the brain surface or specific Regions of Interest (ROIs). Seeing the electrodes in relation to a reconstructed brain surface makes it much easier to verify placement against surgical plans.
3. Data Export Once markers are placed, they can be individually labeled (e.g., "Depth Lead A, Contact 1"). The software then exports these positions as a simple list of indices. This output is designed to be "analysis-ready," allowing researchers to pipe the coordinates directly into their statistical or signal-processing pipelines.
Technical Foundation
I chose Python for this project because of its strength in the scientific community. By leveraging a few key libraries, I was able to keep the tool lightweight but powerful:
-
Nibabel: For robust handling of NIfTI (.nii) files.
-
NumPy: To handle the heavy lifting of coordinate mathematics.
-
VTK: For the 3D engine.
-
GTK+: For a clean, functional user interface.
Open Source and Integration
PyLocator was designed to fit into a larger neuroimaging ecosystem. It plays well with FSL, particularly when working with images that have been normalized to the MNI152 standard template.
By releasing it under a BSD-style license, my goal was to provide a tool that others could not only use but also adapt. While the project began as a solution to a specific research need, it stands as a testament to how open-source tools can simplify the complex spatial problems we face in modern neuroscience.
The source code and documentation for PyLocator remain available for those looking to integrate 3D electrode localization into their own research workflows.